This guide was written for a previous version of VIVO. While the concepts and process for data ingest are still relevant, the specific steps and queries described here may have changed.
note|There is an earlier version of this guide which has a couple of examples for creating a new data property. Please visit Data Ingest Guide v1.1.
VIVO Data Ingest Guide
VIVO release 1.2
Introduction
Data ingest refers to any process of loading existing data into VIVO other than by direct interaction with VIVO's content editing interfaces. Typically this involves downloading or exporting data of interest from an online database or a local system of record.
Data ingest is not all automated – in fact, the current ingest tools involve working through a number of steps from original source data files to the appearance of new data in VIVO. The process requires some understanding of semantic web data modeling and some training, especially until the development, implementation, and user support teams have had more time to work on improving the documentation and the relevant parts of the interface and underlying source code.
VIVO Data Types
VIVO includes basic tools to read CSV (comma-separated values) files typically generated from an Excel file or database report. Normally these files include one row of data per record (e.g., a person or publication) and represent the fields or properties associated with each record in separate columns within the row, much as the values appear in a spreadsheet. The most common pattern of loading CSV files involves one CSV file per type of data to be loaded.
VIVO can also read and convert well-formed XML files from XML to RDF, or Resource Description Framework (http://www.w3.org/RDF/). One way of writing RDF is RDF/XML, and VIVO can use a supplied XSL or Extensible Stylesheet Language transformations (http://www.w3.org/Style/XSL/) to attempt to convert XML to RDF directly. Naturally this success of conversion depends heavily on the structure and consistency of the XML data.
XML files do not store data in a row and column format, but as nested sets of data elements, some of which represent properties or fields analogous to columns. Other elements, however, represent the equivalent of records that may become separate individuals when loaded into VIVO. An XML file of publications, for example, would most likely contain within each block of XML representing one publication a block of one or more author entries. VIVO's ingest process must attempt to pull the authors out of the XML file's publication records and then relate authors and publications.
Mapping Ontologies to other Ontologies
When VIVO's ingest tools read in a CSV or XML file, the data read from the file are stored in the VIVO database as an extra "model," or secondary database managed by the Jena semantic web libraries underlying VIVO.
The next step is to link imported data sets using information stored with the source data. If an object not previously in VIVO has been found, a new record (individual) must be created in VIVO using the CONSTRUCT form of query in the SPARQL query language for RDF.
The query looks at the list of imported data and inserts statements creating a new individual wherever no match to an existing individual is found. The query of imported data is expressed using the ontology of the import; the CONSTRUCT statements use the class and property names of VIVO ontology. This accomplishes the mapping between the source ontology and the VIVO ontology.
When populating VIVO for the first time, all the data from a dataset can be added from the imported Jena model to VIVO by a CONSTRUCT query that again translates from the RDF in the imported model to the RDF in VIVO. When a dataset has already been mapped to data and there's a likelihood that the new data being imported may already exist in VIVO, the process becomes more complicated due to the need to match each prospective import against existing data.
Data Ingest Example
The first time through an ingest process is a slow process of evaluating the source data, deciding what cleanup will be needed, and working through each step. The following example details each step in the process and how the VIVO software supports that step. As the process expands to include requirements to match against existing data, additional steps must be introduced to test for matches and perform different actions, usually as successive steps identified through different queries.
For this example, a sample set of data for people, their positions, and the organizations at which their positions reside has been provided at http://sourceforge.net/projects/vivo/files/Data%20Ingest/. The guide gives instructions on how to ingest the data, construct the data to the ontology, and load the data into an RDF format. Upon completion linked data in VIVO will include people to positions and the positions to organizations.
Process:
- Create a local ontology
- Create workspace models for ingesting and constructing data
- Pull external data file into RDF
- Map tabular data into the ontology format
- Construct the ingested entities using the map of properties
- Load data to the current web model
Step 1: Create a Local Ontology
When VIVO is first installed it comes with eight ontologies preloaded. It is recommended that each institution creates a local ontology to accommodate the unique needs of the institute’s data and data source.
For the demonstration, we are going to create data properties within a local ontology to accommodate data source information. We will start by creating the local ontology, and then add a data property for a person and an organization’s unique identifier.
Create Local Ontology
- Select the 'Site Admin' link in top right corner to return to the Site Administration page
- Select 'Ontology List' in the Ontology Editor section
- Select 'Add new ontology'
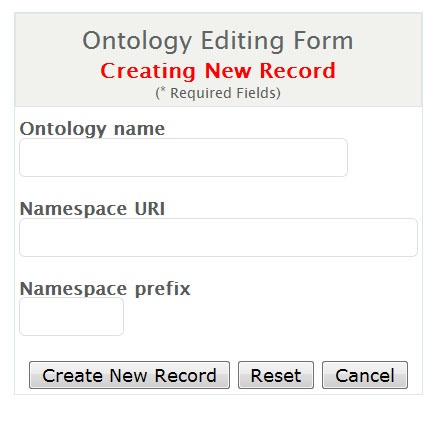
- Input information as noted in the figure below and select 'Create New Record'
Create a New Data Property
- Select the newly created ontology
- Select 'Datatype Properties Defined in This Namespace'
- Select 'Add new data property'
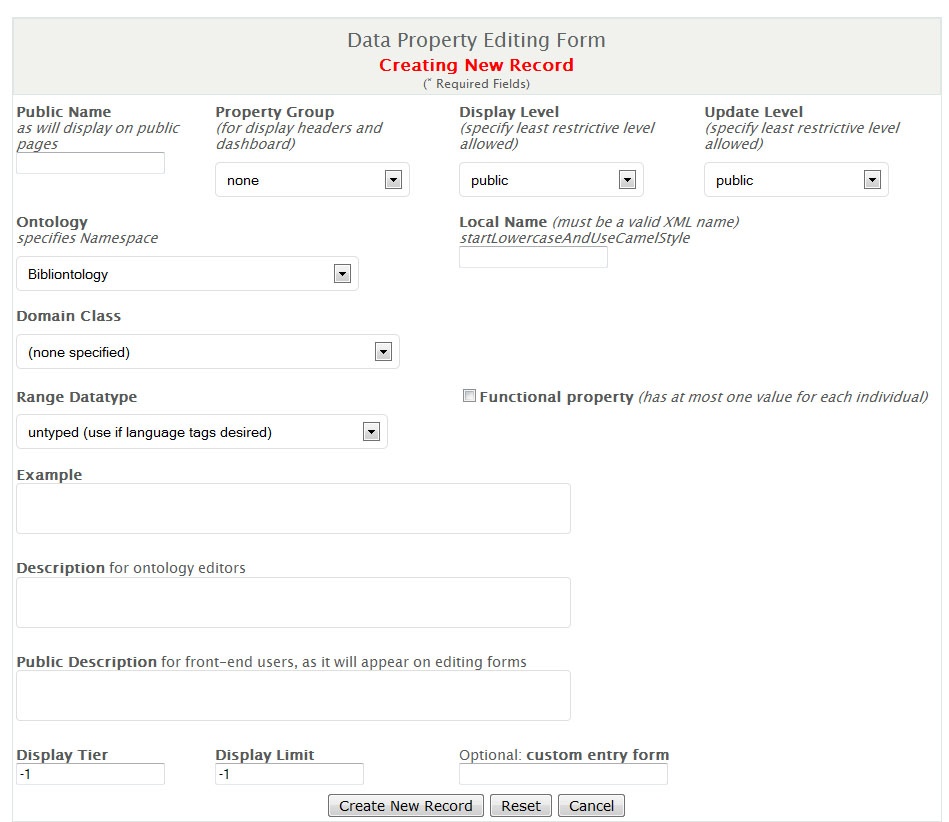
- Add the information as follows and select 'Create New Record'
- Repeat steps 1 through 4 for an Organization ID
Within the main model (webapps) we can now maintain the unique four digit code that links a person across a variety of data sources. In the main model (webapps) we can also maintain the organization ID, the main identifier for budgetary unit or responsibility center associated with the “University of VIVO”.
Data Ingest Menu
Highlighted in red are the three Ingest Menu options we will be using for this demonstration.
Step 2: Create Workspace Models
Working with VIVO requires pulling data into a model and transforming the data to create the proper assertions. It is best to create a couple of models, at least an ingest model and construction model, to use as you run through the data ingest.
- To perform this step, select "Ingest Tools" from the Advanced Tools Menu
- Select "Manage Jena Models"
- Select "Create Model" then type in a name for your model (it can be anything)
In this example we will create two new models named as follows:
- csv-import-model
- csv-construct-model
The Manage Jena Models page is very useful for other steps. You can attach the model to the current webapp to see the created entities. You can output the model to see the work that is created. You can clear out a model to wipe the slate clean and start over.
Step 3: Pull External Data File into RDF
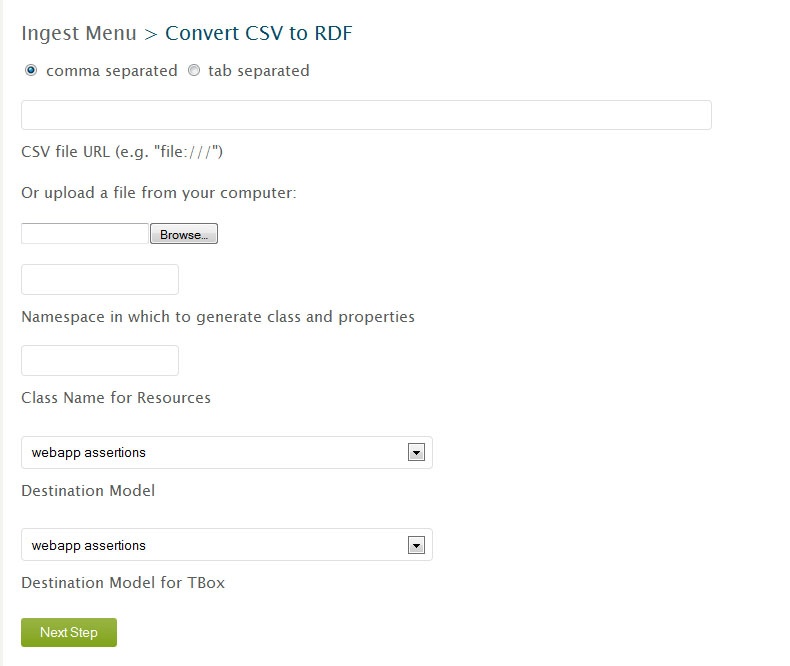
Ingesting data from external sources can be found under the ingest menu, "Convert CSV to RDF". The first box "CSV file URL" is for your CSV file. Your CSV file will need to be in a public web location or on the local server hosting the VIVO application. When the file is on the same server as VIVO you can type file:///_filelocation.
CSV file URL
The three files needed for this workshop can also be found on our public web location:
http://sourceforge.net/projects/vivo/files/Data%20Ingest/people.csv/download
http://sourceforge.net/projects/vivo/files/Data%20Ingest/organization.csv/download
http://sourceforge.net/projects/vivo/files/Data%20Ingest/position.csv/download
Namespace for Resources
The namespace for the entities you create will follow the entity in this example. So the first name space should follow the format of your ontology. The default virtual appliance setting is *
http://localhost/vivo/
*, however most institutions follow the format *
http://vivo.institution.edu/
*.
Namespace for Classes and Properties
The namespace for the properties does not follow the created entity since you will shift the properties from the format they come in into the ontologies format, so this name space can be a temporary namespace. However, it will not hurt your program to also use the same default namespace, *
http://localhost/vivo/
*.
Class Name
The class name is also a temporary value for this example. This value does not follow the created entity since you will shift the properties from the format they come in into the ontologies format. For this example, the suggested class names are “ws_ppl”, “ws_org”, and “ws_post”.
Destination Models
The "Destination Model" and "Destination Model for TBox" should list the model to ingest into. This is where we select one of those 'workspace' models we created earlier, "csv-import-model" for example.
After clicking convert we can check our RDF conversion through two methods, the first method being the quickest.
- From the Ingest Menu, select the 'Manage Jena Models' and then select the ingest model's 'output model.' This is something you can open with WordPad and see the created triples for your CSV file.
- Also from the Manage Jena Model we can attach the ingest model to the current webapp. Then we can go back to the Site Admin, select Class Hierarchy link, select 'All Classes' and navigate to the Class Name you created (individual, for example).
Step 4: Map Tabular Data onto Ontology
To create a SPARQL Query to construct the correct entities we need gather the URI's for the properties, classes, and associations that we are going to make, and the URI's for the ingested data.
Ingested Data URI's
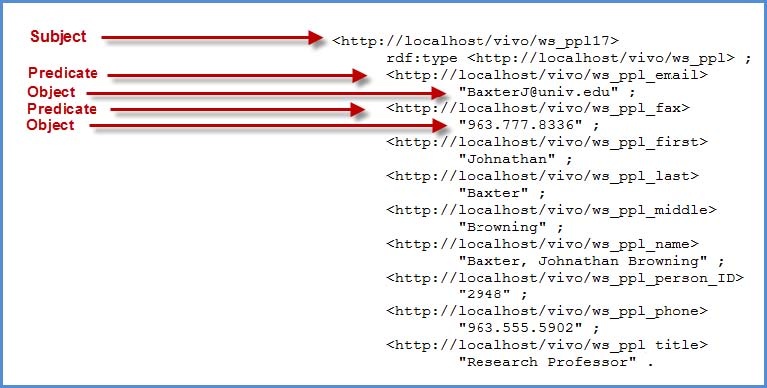
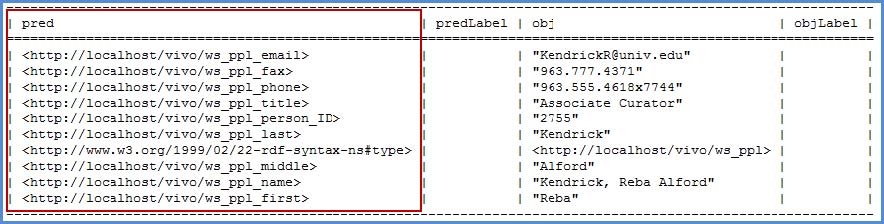
The easiest method to attaining the URI's that were created for each of the columns in your csv file, is to open the “output model” rdf data as described above. A description of the format is described in the figure below. The predicates are the properties we need for our SPARQL Query.
An alternate method that displays the results in an easier to read output is shown below.
- Click ‘attach to webapp’ to attach the model of the ingested data
- Navigate to the root classes and locate the class name you created
- Click Site Admin, then ‘Class hierarchy’ in the Ontology Editor section
- Scroll down and select the class ws_ppl
- Select 'Show all individuals in this class' from the Class Control Panel
- Select any individual
- Select 'Raw Statements with this Resource as Subject'
To retrieve the URI's for the ingested properties we selected "Raw Statements with this Resource as Subject". This will output in text format the predicates and objects where the individual is a resource. These predicates are the properties we need for our SPARQL Query. Copy them to a notepad file so we can retrieve them later.
Ontology Properties
If you already have individuals in your installation you can use the same four methods above to find the predicates to be constructed. Otherwise simply create a new entity and click submit at the bottom of the page. Now select 'Raw Statements with this Resource as Subject' to see the predicates.
- Select Site Admin at the top right hand corner of the page
- Select a Faculty person type in the Data Input section
- Select 'Add individual of this class'
- Name the faculty member and select create
- Select ‘Edit this individual’
- Select ‘Edit this Individual’ (again)
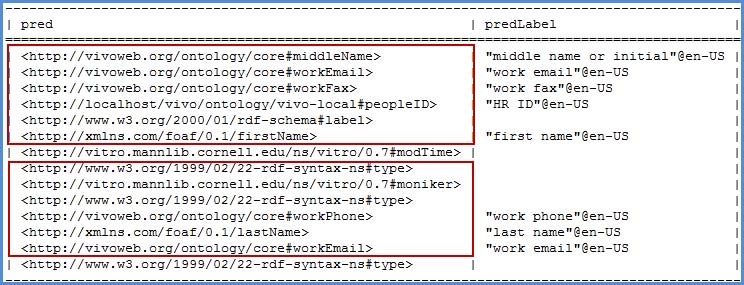
- Populate the fields that are related to the data in the csv file (middle name, work email, work fax)
- Select 'Raw Statements with this Resource as Subject'
- Copy the predicates needed for our SPARQL Query to a notepad for later use
An alternate method that displays the results is to navigate to the ontology lists of Class hierarchy, Data Properties and Object Properties to find the various URI's we need. Opening a class or property from the list will give an overview of the item. At the bottom of that overview is the URI needed to associate the ingested data with the ontologies used in VIVO. Copy the URI for each data property contained within your CSV file.
Step 5: Construct the Ingested Entities
When writing a SPARQL Query ensure that the URI's are constructed from the CSV to RDF and the properties we want the data to be placed into.
Click on 'Execute SPARQL CONSTRUCT'
Constructing new entities in SPARQL starts with the basic frame:
Construct
{
}
Where
{
}
We then need to create statements that will create triples in VIVO in the basic format (subject predicate object). The period ‘.’ separates triples from each other. Variables are represented by the question mark ‘? ‘ before the name of the variable.
Example for people.csv:
Construct
{
?person <http://www.w3.org/1999/02/22-rdf-syntax-ns#type> <http://vivoweb.org/ontology/core#FacultyMember> .<br/>
?person <http://www.w3.org/2000/01/rdf-schema#label> ?fullname .<br/>
?person <http://xmlns.com/foaf/0.1/firstName> ?first .<br/>
?person <http://vivoweb.org/ontology/core#middleName> ?middle .<br/>
?person <http://xmlns.com/foaf/0.1/lastName> ?last .<br/>
?person <http://vitro.mannlib.cornell.edu/ns/vitro/0.7#moniker> ?title .<br/>
?person <http://vivoweb.org/ontology/core#workPhone> ?phone .<br/>
?person <http://vivoweb.org/ontology/core#workFax> ?fax .<br/>
?person <http://vivoweb.org/ontology/core#workEmail> ?email .<br/>
?person <http://vivoweb.org/ontology/vivo-local#peopleID> ?HRID .<br/>
}
Where
{
?person <http://localhost/vivo/ws_ppl_name> ?fullname .<br/>
?person <http://localhost/vivo/ws_ppl_first> ?first .<br/>
optional { ?person <http://localhost/vivo/ws_ppl_middle> ?middle . }<br/>
?person <http://localhost/vivo/ws_ppl_last> ?last .<br/>
?person <http://localhost/vivo/ws_ppl_title> ?title .<br/>
?person <http://localhost/vivo/ws_ppl_phone> ?phone .<br/>
?person <http://localhost/vivo/ws_ppl_fax> ?fax .<br/>
?person <http://localhost/vivo/ws_ppl_email> ?email .<br/>
?person <http://localhost/vivo/ws_ppl_person_ID> ?HRID .<br/>
}
Next:
- Select Source Model (‘csv-ingest’)
- Select Destination Model (‘csv-construct’)
- Select "Execute CONSTRUCT"
- Upon completion, the system will report ‘375 statements CONSTRUCTed’.
Some interesting things to note:
<http://www.w3.org/1999/02/22-rdf-syntax-ns#type> is used to associate entities with classes. You are basically making the assertion John Doe is a Faculty Member.
<http://www.w3.org/2000/01/rdf-schema#label> is the predicate that is displayed when an entity is returned from a search, and at the top of their page. In many situations, this is the name of the entity.
<http://vitro.mannlib.cornell.edu/ns/vitro/0.7#moniker> is the predicate that serves as a sub-label, think of it as the title of an entity. It is also displayed from a search, under the label predicate. A person would be Jane Doe with a moniker of Special Title Professor. A Department might be Donor Name College of Fine Arts with a moniker that describes the schools that make up the college.
Step 6: Load to Webapp
The live webapp that is indexed does not allow for models to just be attached. Attaching models works well for seeing what we have constructed or ingested or smushed, but those models are lost when Tomcat refreshes.
The final step is to output the final model and add it into the current model
- From the Ingest Menu, select "Manage Jena Models"
- Select "output model" for the desired construct model
- Save the resulting file
- Navigate back to the Site Administration page
- Select Add/Remove RDF
- Browse to the file previously saved
- Select N3 as import format*
- Confirmation should state ‘Added RDF from file people_rdf. Added 415 statements.’
- The output engine uses N3, not RDF/XML. This is important to note when adding the ‘output mode’ RDF data into the webapp. RDF/XML is the default setting for the drop down list as most ontologies are written in RDF/XML.
The ingested data should now display in both the index and the search results. It is part of the main webapp and will be retained upon a tomcat restart. The founding steps of data ingest have been completed.
Process Review
- Created a local ontology
- Created workspace models for ingesting and constructing data
- Pulled external data file into RDF
- Mapped tabular data into the ontology format
- Constructed the ingested entities using the map of properties
- Loaded data to the current web model
Included with this guide are example CSV data files, SPARQL construct queries, and the resulting RDF data files for people, organizations, and positions. Please consult these files when performing the data ingest of organizations and positions.
Note: To perform the construct for ‘position to people’ and ‘position to organization’ noted in the appendix, both the position csv and the associated file (people or organization) need to be loaded into the same model. Additional information on this process will be included in future releases of this guide.
Appendix A: SPARQL Queries
People Construct:
Construct
{
?person <http://www.w3.org/1999/02/22-rdf-syntax-ns#type> <http://vivoweb.org/ontology/core#FacultyMember> .<br/>
?person <http://www.w3.org/2000/01/rdf-schema#label> ?fullname .<br/>
?person <http://xmlns.com/foaf/0.1/firstName> ?first .<br/>
?person <http://vivoweb.org/ontology/core#middleName> ?middle .<br/>
?person <http://xmlns.com/foaf/0.1/lastName> ?last .<br/>
?person <http://vitro.mannlib.cornell.edu/ns/vitro/0.7#moniker> ?title .<br/>
?person <http://vivoweb.org/ontology/core#workPhone> ?phone .<br/>
?person <http://vivoweb.org/ontology/core#workFax> ?fax .<br/>
?person <http://vivoweb.org/ontology/core#workEmail> ?email .<br/>
?person <http://localhost/vivo/ontology/vivo-local#peopleID> ?hrid .<br/>
}
Where
{
?person <http://localhost/vivo/ws_ppl_name> ?fullname .<br/>
?person <http://localhost/vivo/ws_ppl_first> ?first .<br/>
optional { ?person <http://localhost/vivo/ws_ppl_middle> ?middle . }
?person <http://localhost/vivo/ws_ppl_last> ?last .<br/>
?person <http://localhost/vivo/ws_ppl_title> ?title .<br/>
?person <http://localhost/vivo/ws_ppl_phone> ?phone .<br/>
?person <http://localhost/vivo/ws_ppl_fax> ?fax .<br/>
?person <http://localhost/vivo/ws_ppl_email> ?email .<br/>
?person <http://localhost/vivo/ws_ppl_person_ID> ?hrid .<br/>
}
Organization Construct:
Construct
{
?org <http://www.w3.org/1999/02/22-rdf-syntax-ns#type> <http://xmlns.com/foaf/0.1/Organization> .<br/>
?org <http://localhost/vivo/ontology/vivo-local#orgID> ?deptID .<br/>
?org <http://www.w3.org/2000/01/rdf-schema#label> ?name .<br/>
}
Where
{
?org <http://localhost/vivo/ws_org_org_ID> ?deptID .
?org <http://localhost/vivo/ws_org_org_name> ?name .
}
Position to People Construct:
Construct
{
?position <http://www.w3.org/1999/02/22-rdf-syntax-ns#type> <http://vivoweb.org/ontology/core#FacultyPosition> .<br/>
?position <http://vivoweb.org/ontology/core#startYear> ?year .<br/>
?position <http://www.w3.org/2000/01/rdf-schema#label> ?title .<br/>
?position <http://vivoweb.org/ontology/core#titleOrRole> ?title .<br/>
?position <http://vivoweb.org/ontology/core#positionForPerson> ?person .<br/>
?person <http://vivoweb.org/ontology/core#personInPosition> ?position .<br/>
}
Where
{
?position <http://localhost/vivo/ws_post_department_ID> ?orgID .<br/>
?position <http://localhost/vivo/ws_post_start_date> ?year .<br/>
?position <http://localhost/vivo/ws_post_job_title> ?title .<br/>
?position <http://localhost/vivo/ws_post_person_ID> ?posthrid .<br/>
?person <http://localhost/vivo/ws_ppl_person_ID> ?perhrid .<br/>
FILTER((?posthrid)=(?perhrid))
}
Position to Organization Construct:
Construct
{
?position <http://www.w3.org/1999/02/22-rdf-syntax-ns#type> <http://vivoweb.org/ontology/core#FacultyPosition> .<br/>
?position <http://vivoweb.org/ontology/core#startYear> ?year .<br/>
?position <http://www.w3.org/2000/01/rdf-schema#label> ?title .<br/>
?position <http://vivoweb.org/ontology/core#titleOrRole> ?title .<br/>
?org <http://vivoweb.org/ontology/core#organizationForPosition> ?position .<br/>
?position <http://vivoweb.org/ontology/core#positionInOrganization> ?org .<br/>
}
Where
{
?position <http://localhost/vivo/ws_post_start_date> ?year .<br/>
?position <http://localhost/vivo/ws_post_job_title> ?title .<br/>
?position <http://localhost/vivo/ws_post_department_ID> ?postOrgID .<br/>
?org <http://localhost/vivo/ws_org_org_ID> ?orgID .<br/>
FILTER((?postOrgID)=(?orgID))
}